Tuesday, February 18, 2014 From rOpenSci (https://ropensci.org/blog/2014/02/18/antweb/). Except where otherwise noted, content on this site is licensed under the CC-BY license.
This post was updated on August 20, 2014, with
AntWebversion0.7.2.99. Please install an updated version to make sure the code works.

Data on more than 10,000 species of ants recorded worldwide are available through from California Academy of Sciences’ AntWeb, a repository that boasts a wealth of natural history data, digital images, and specimen records on ant species from a large community of museum curators.
Digging through some of the earliest announcements of AntWeb, I came across a Nature News piece titled “Mashups mix data into global service” from January 2006. The article contains this great quote from Roderic Page “If you could pool data from every museum or lab in the world, you could do amazing things”. The article also says “So far, only researchers with advanced programming skills, working in fields organized enough to have data online and tagged appropriately, have been able to do this.” In many ways this really is motivation for why we develop interfaces to these rich data repositories. Our express intent is to facilitate researchers explore amazing opportunities that lie within such data by lowering techinical barriers to use. Right on the heels of our most recent package (ecoengine), we are now happy to first release of an interface to AntWeb.
A stable version of our R package AntWeb is now available from CRAN. The API currently does not require a key for access but larger requests will be throttled on the server side. It is worth noting that much of these same data are also ported through the Global Biodiversity Information Facility and accessible through our gbif package. This package provides a more direct interface to more of the ant specific natural history data.
🔗 Installing the package
A stable version of the package (0.7) is now available on CRAN.
install.packages("AntWeb")
or you can install the latest development version (the master branch is also always stable & deployable and most up-to-date. Current version is 0.5.3 at the time of this writing).
library(devtools)
install_github("ropensci/AntWeb")
🔗 Searching through the database
As with most of our packages, there are several ways to search through an API. In the case of AntWeb, you can search by a genus or full species name or by other taxonomic ranks like sub-phylum.
Data on ants
To obtain data on any taxonomic group, you can make a request using the aw_data() function. It’s possible to search easily by a taxonomic rank (e.g. a genus) or by passing a complete scientific name.
Searching by Genus
library(AntWeb)
# To get data on an ant genus found widely through Central and South America
data_genus_only <- aw_data(genus = "acanthognathus")
430 results available for query.
leaf_cutter_ants <- aw_data(genus = "acromyrmex")
713 results available for query.
unique(leaf_cutter_ants$data$scientific_name)
[1] "acromyrmex versicolor" "acromyrmex striatus"
[3] "acromyrmex balzani" "acromyrmex coronatus"
[5] "acromyrmex crassispinus" "acromyrmex heyeri"
[7] "acromyrmex lundii" "acromyrmex fracticornis"
[9] "acromyrmex niger" "acromyrmex nigrosetosus"
[11] "acromyrmex rugosus" "acromyrmex subterraneus"
[13] "acromyrmex alw01" "acromyrmex alw02"
[15] "acromyrmex alw03" "acromyrmex alw04"
[17] "acromyrmex octospinosus" "acromyrmex lobicornis"
[19] "acromyrmex silvestrii" "acromyrmex landolti"
[21] "acromyrmex ambiguus" "acromyrmex hystrix"
[23] "acromyrmex laticeps" "acromyrmex indet"
[25] "acromyrmex echinatior" "acromyrmex volcanus"
[27] "acromyrmex disciger" "acromyrmex aspersus"
[29] "acromyrmex pubescens" "acromyrmex moelleri"
[31] "acromyrmex evenkul" "acromyrmex hispidus"
[33] "acromyrmex nobilis" "acromyrmex pulvereus"
[35] "acromyrmex lundi"
Searching by species
# You can request data on any particular species
acanthognathus_df <- aw_data(scientific_name = "acanthognathus brevicornis")
2 results available for query.
head(acanthognathus_df)
$count
[1] 2
$call
$call$genus
[1] "acanthognathus"
$call$species
[1] "brevicornis"
$data
url
1 http://antweb.org/api/v2/?occurrenceId=CAS:ANTWEB:casent0280684
2 http://antweb.org/api/v2/?occurrenceId=CAS:ANTWEB:casent0637708
catalogNumber family subfamily genus specificEpithet
1 casent0280684 formicidae myrmicinae Acanthognathus brevicornis
2 casent0637708 formicidae myrmicinae Acanthognathus brevicornis
scientific_name typeStatus stateProvince country
1 acanthognathus brevicornis Colombia
2 acanthognathus brevicornis
dateIdentified habitat minimumElevationInMeters
1 NA
2 2013-09-12 NA
# You can also limit queries to observation records that have been geoferenced
acanthognathus_df_geo <- aw_data(genus = "acanthognathus", species = "brevicornis", georeferenced = TRUE)
It’s also possible to search for records around any location by specifying a search radius.
data_by_loc <- aw_coords(coord = "37.76,-122.45", r = 2)
# This will search for data on a 2 km radius around that latitude/longitude
Image data
Most specimens in the database have images associated with them. These include high, medium, and low resolution version of the head, dorsal side, full profile, and the specimen label. For example we can retrieve data on a specimen of Ecitoninaeeciton burchellii with the following call:
# Data and images for Ecitoninaeeciton burchellii
eb <- aw_code(occurrenceid ="CAS:ANTWEB:casent0003205")
eb$image_data$high[[2]]
NULL
If you’re primarily interested in ant images and would like to keep up with recent additions to the database, you can also use the aw_images function. This function takes two arguments: since, the number of days to search backward, and a type. Possible options for type are h for head, d for dorsal, p for profile, and l for label. If a type is not specified, all available images are retrieved.
# Retrieve only dorsal images for the last five days
head(aw_images(since = 5, img_type = "d"))
high
1 https://www.antweb.org/images/casent0914000/casent0914000_d_1_high.jpg
2 https://www.antweb.org/images/antweb1008677/antweb1008677_d_1_high.jpg
3 https://www.antweb.org/images/casent0906977/casent0906977_d_1_high.jpg
4 https://www.antweb.org/images/casent0914012/casent0914012_d_1_high.jpg
5 https://www.antweb.org/images/antweb1008691/antweb1008691_d_1_high.jpg
6 https://www.antweb.org/images/casent0906997/casent0906997_d_1_high.jpg
med
1 https://www.antweb.org/images/casent0914000/casent0914000_d_1_low.jpg
2 https://www.antweb.org/images/antweb1008677/antweb1008677_d_1_low.jpg
3 https://www.antweb.org/images/casent0906977/casent0906977_d_1_low.jpg
4 https://www.antweb.org/images/casent0914012/casent0914012_d_1_low.jpg
5 https://www.antweb.org/images/antweb1008691/antweb1008691_d_1_low.jpg
6 https://www.antweb.org/images/casent0906997/casent0906997_d_1_low.jpg
low
1 https://www.antweb.org/images/casent0914000/casent0914000_d_1_med.jpg
2 https://www.antweb.org/images/antweb1008677/antweb1008677_d_1_med.jpg
3 https://www.antweb.org/images/casent0906977/casent0906977_d_1_med.jpg
4 https://www.antweb.org/images/casent0914012/casent0914012_d_1_med.jpg
5 https://www.antweb.org/images/antweb1008691/antweb1008691_d_1_med.jpg
6 https://www.antweb.org/images/casent0906997/casent0906997_d_1_med.jpg
thumbnail
1 https://www.antweb.org/images/casent0914000/casent0914000_d_1_thumbview.jpg
2 https://www.antweb.org/images/antweb1008677/antweb1008677_d_1_thumbview.jpg
3 https://www.antweb.org/images/casent0906977/casent0906977_d_1_thumbview.jpg
4 https://www.antweb.org/images/casent0914012/casent0914012_d_1_thumbview.jpg
5 https://www.antweb.org/images/antweb1008691/antweb1008691_d_1_thumbview.jpg
6 https://www.antweb.org/images/casent0906997/casent0906997_d_1_thumbview.jpg
img_type catalog_id
1 d https://www.antweb.org/api/v2/?catalogNumber=casent0914000
2 d https://www.antweb.org/api/v2/?catalogNumber=antweb1008677
3 d https://www.antweb.org/api/v2/?catalogNumber=casent0906977
4 d https://www.antweb.org/api/v2/?catalogNumber=casent0914012
5 d https://www.antweb.org/api/v2/?catalogNumber=antweb1008691
6 d https://www.antweb.org/api/v2/?catalogNumber=casent0906997
upload_date
1 2014-08-15 15:11:14
2 2014-08-20 13:52:13
3 2014-08-15 15:11:14
4 2014-08-15 15:11:15
5 2014-08-20 13:52:13
6 2014-08-15 15:11:15
It’s also possible to retrieve unique lists of any taxonomic rank using the aw_unique function.
subfamily_list <- aw_distinct(rank = "subfamily")
subfamily_list
[Total results on the server]: 67
[Args]:
rank = subfamily
limit = 1000
[Dataset]: [67 x 1]
[Data preview] :
[1] apidae bethylidae
67 Levels: aenictinae agroecomyrmecinae amblyoponinae ... xylocopinae
genus_list <- aw_distinct(rank = "genus")
genus_list
[Total results on the server]: 490
[Args]:
rank = genus
limit = 1000
[Dataset]: [490 x 1]
[Data preview] :
[1] Aenictinae Amblyoponinae
490 Levels: Acanthognathus Acanthomyrmex Acanthoponera ... Zigrasimecia
species_list <- aw_distinct(rank = "species")
species_list
[Total results on the server]: 10547
[Args]:
rank = species
limit = 1000
[Dataset]: [1000 x 1]
[Data preview] :
[1] basicerotini indet
1000 Levels: a abbreviata abdelazizi abdera abdita abditivata ... orizabanum_complex
If you work with existing specimens, you can also query directly by a specimen ID.
(data_by_code <- aw_code(catalogNumber="inb0003695883"))
[Total results on the server]: 1
[Args]:
catalogNumber = inb0003695883
[Dataset]: [1 x 16]
[Data preview] :
specimens.url
1 http://antweb.org/api/v2/?occurrenceId=CAS:ANTWEB:inb0003695883
NA <NA>
specimens.catalogNumber specimens.family specimens.subfamily
1 inb0003695883 formicidae myrmicinae
NA <NA> <NA> <NA>
specimens.genus specimens.specificEpithet specimens.scientific_name
1 Acanthognathus teledectus acanthognathus teledectus
NA <NA> <NA> <NA>
specimens.typeStatus specimens.stateProvince specimens.country
1 Heredia Costa Rica
NA <NA> <NA> <NA>
specimens.dateIdentified specimens.habitat
1 2006-11-02 mature wet forest ex sifted leaf litter
NA <NA> <NA>
specimens.minimumElevationInMeters specimens.geojson.type
1 50 point
NA <NA> <NA>
decimal_latitude decimal_longitude
1 10.413477 -84.023636
NA <NA> <NA>
# This will return a list with a metadata data.frame and a image data.frame
If you have a multiple specimen IDs, as is often the case when working with research data, you can get data on all of them at the same time. The function automatically retuns NULL values when no data are found and you can have these removed using plyr::compact (this happens automatically when you use a function call like ldply.)
specimens <- c("casent0908629", "casent0908650", "casent0908637")
results <- lapply(specimens, function(x) aw_code(x))
names(results) <- specimens
length(results)
[1] 3
🔗 Mapping ant specimen data
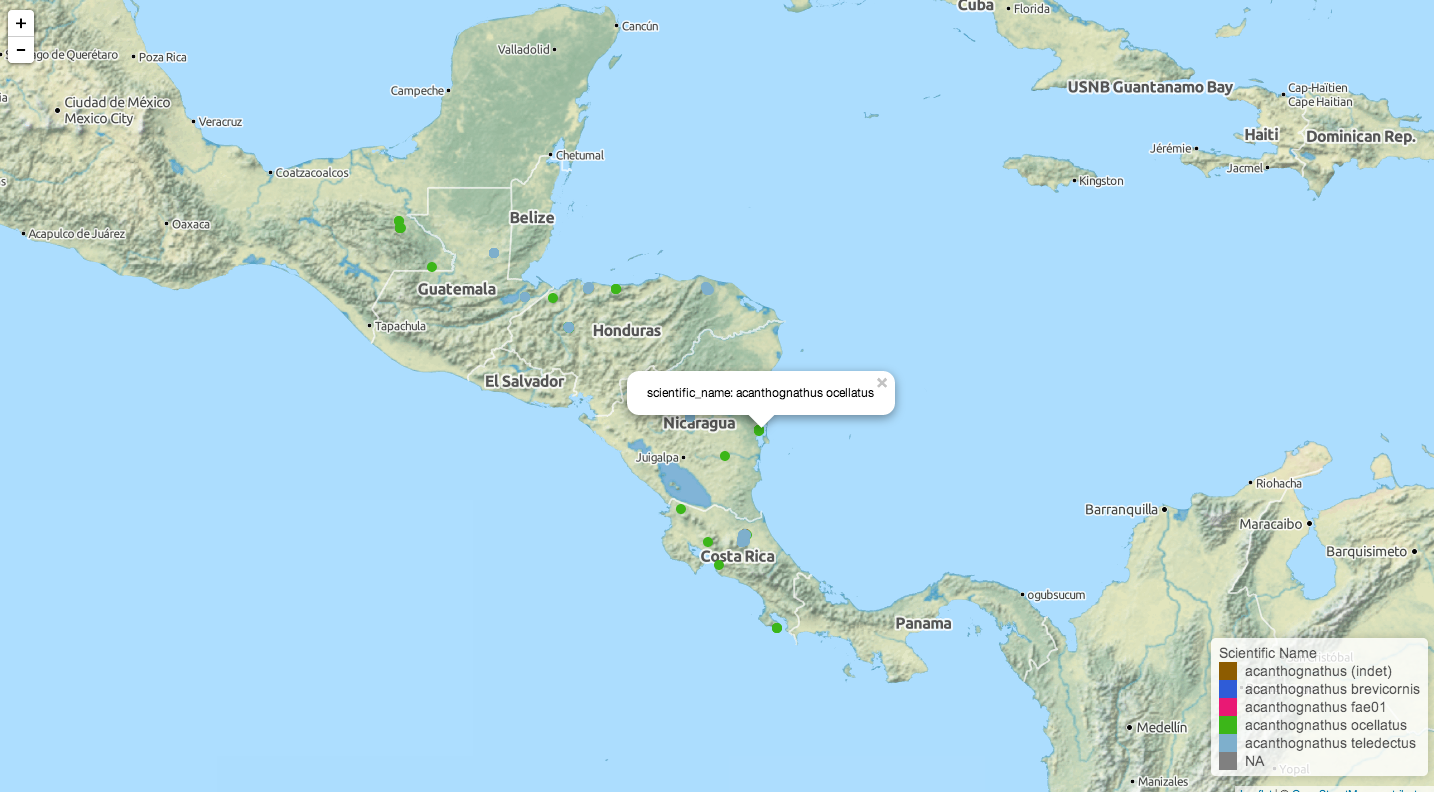
As with the previous ecoengine package, you can also visualize location data for any set of species. Adding georeferenced = TRUE to a data retrieval call will filter out any data points without location information. Once retrieved the data are mapped with the open source Leaflet.js and pushed to your default browser. Maps and associated geoJSON files are also saved to a location specified (or defaults to your /tmp folder). This feature is only available on the development version on GitHub (0.5.2 or greater; see above on how to install) and will be available from CRAN in version 0.6
acd <- aw_data(genus = "acanthognathus")
aw_map(acd)
🔗 Integration with the rest of our biodiversity suite
Our newest package on CRAN, spocc (Species Occurrence Data), currently in review at CRAN, integrates AntWeb among other sources. More details on spocc in our next blog post.
As always please send suggestions, bug reports, and ideas related to the AntWeb R package directly to our repo.

