Tuesday, February 9, 2021 From rOpenSci (https://ropensci.org/blog/2021/02/09/coder/). Except where otherwise noted, content on this site is licensed under the CC-BY license.

A new R-package, coder, has been developed, peer-reviewed by rOpenSci, accepted by CRAN, and published in a paper by the Journal of Open Source Software (JOSS). In this blog post, I will explain why this package might be useful for (epidemiological/medical/health care related) research.
🔗 Clinical mess
Once upon a time, in countries not far from ours, there were MDs and nurses making up funny names for any diseases they encountered. What Dr. A called X would be recognized only as Y by his dear colleague Dr. B. X, however, was also a name used by Dr. C, but then for a completely different condition. It was a mess!
Those times are long gone due to the fascinating story of standardized medical nomenclatures and coding. It all started in the first half of the 19th century and has been well documented in a free on-line book by Iwao Moryama, Ruth Loy and Alastair Robb-Smith.
Today, when visiting the hospital, our (somatic) conditions are most likely recorded in a medical register by some adaptation of the 10th revision of the International Statistical Classification of Diseases and Related Health Problems (ICD-10). This is a global standard for medical coding administrated by the World Health Organization (WHO). Psychiatric conditions might as well be recorded by the Diagnostic and Statistical Manual of Mental Disorders (DSM). Prescribed medical substances are reported by the Anatomical Therapeutic Chemical (ATC) classification system. Surgical procedures performed in the Nordic countries are coded by the Nordic Medico-Statistical Committee’s (NOMESCO’s) Classification of Surgical Procedures (NCSP). All combined, there are several hundred thousands of different codes quantifying every possible interaction with health care. Although the mess has significantly decreased, the world of medical coding is still rather messy! Not only are there different versions of ICD (currently with ICD-11 in the pipe). There are also yearly revisions of the currently used version, as well as clinical versions, oncological versions, and several national implementations. All this might be well motivated from a clinical and/or administrative view-point, but it does make things difficult for researchers trying to pool data from different sources, countries and periods.
🔗 Comorbidity
A measure of patient comorbidity is used to summarize the total pre-existing disease burden prior to other diagnosis or events of interest. Such data are often used to control for case-mix confounding in statistical models with various exposures and outcomes. There is no obvious definition of what to consider a comorbid condition, but several suggestions exist. Charlson et al., Elixhauser et al., Sloan et al. and others, have combined individual medical conditions into more holistic categories/diseases. The number of categories might then be summarized by an index value (a weighed sum), as a single comorbidity-measure for each patient. There are several versions of each index, however. To use those indices in practice is also cumbersome since many data sources (large administrative registers) are big - bigger than what would fit in the random access memory (RAM) of a standard computer.
🔗 R packages
There are some excellent R-packages to help with both the Charlson and Elixhauser comorbidity indices. comorbidity by Alessandro Gasparini and icd by Jack O. Wasey and Michael Lang are very well documented with efficient implementations. To keep up with all different and newly proposed coding versions seems like a daunting, if not impossible task, however. Therefore, the coder package offers an alternative, more flexible, implementation. It is not hard-wired to Charlson or Elixhauser, although capabilities for those indices are included as well. data.table is used internally for computational efficiency, while the application programming interface (API) is more inspired by tidyverse.
🔗 Regular expressions
In coder, all relevant codes (from ICD, ATC, NCSP or whatever) are represented in a compact way by their corresponding regular expressions. Peptic ulcer disease for example is recognized as an important comorbidity by the Elixhauser classification. It might be coded by ICD-10 as “K25.x–K28.x” (see table 2 here). This format is well understood by humans; “-” indicates an interval and “x” acts like a wild card for any additional alphanumerical characters. This might be translated into a code list (ignoring the dots): K257, K259, K267, K269, K277, K279, K287, K289. But why those codes only? Why not K250 or K289A? Well, not all codes are used in practice. This list is just an example corresponding to codes defined in the clinical implementation of ICD-10 (ICD-10-CM) used in 2020 in the USA. The list might look different with data from the Swedish version of ICD-10 (ICD-10-SE) recorded in 2021. Hence, the use of regular expressions will relax the need of a complete and version-specific code list.
A regular expression for peptic ulcer disease might be ^K2[5-8][79]$ where ^ ($) marks the beginning (end) of the character string, where alternative digits are written within brackets, and where the generic wild card x has been limited to either 7 or 9 as used in clinical practice (see ?regex in R for details).
To use regular expressions is also more computationally efficient compared to a traditional approach where each individual code might be compared by exact matches to any “word” (cell) in a long dictionary (vector) of individual codes. Hence, it is easy to realize that the code K257 can not be found in a dictionary limited to words with initials L to Z. Thus, you do not need to compare the code K257 to each and every entry in that list to see if you get a match or not.
Such computational efficiency is crucial if working with large populations (as from a national patient register spanning several decades et cetera).
By default, there are 413 of those regular expressions included in the coder package. They cover all conditions recognized by Charlson, Elixhauser, RxRisk V, the comorbidity-polypharmacy score (CPS), as well as for some diagnose specific adverse events classifications after hip or knee replacement. Charlson, Elixhauser and Rx Risk V have 6, 5 and 3 alternative regular expressions for each condition. Peptic ulcer disease for example can also be recognized from 1) 53([1-4][79]0)|V1271; 2) 53([1-4]([4-69]1|7)); or 3) 53[1-4][79] based on three different versions of ICD-9-CM. Additional versions (think of the upcoming ICD-11) are relatively easy to implement by using a dedicated classcodes object (an enhanced data frame/tibble with alternative regular expressions in each column).
🔗 Interpretation
Although classcodes objects are relatively easy to implement, the interpretation of a regular expression might be hard to grasp intuitively.
Or what about hypertension according to Pratt’s version of the Rx Risk V classification based on ATC codes: C0(2(A(B[0-9]{2}|C0[0-5])|DB(0[2-9]|[1-9][0-9]))|3((A[A-Z][0-9]{2}|BA(0[0-9]|1[01]))|DB(01|99)|C([AB][0-9]{2}|C0[01])|EA01)|9(BA0[2-9]|DA0[2-8]|C[A-X][0-9]{2}))?
Fortunately, there are several functions in the coder package to aid such interpretation:
visualize()will open the default web browser to show a structural diagram based on the JavaScript Regular Expression Visualizer.summary()relies on the decoder package to decode the meaning of a regular expression in the content of a specified code version. For example the ICD-10 specification of a dislocation after total hip arthroplasty (^(M24(3|4F?)|S730|T933)$) would expand to 5 individual codes according to ICD-10-SE: M243, M244, M244F, S730, T933. None of those would be recognized by ICD-10-CM, however. A longer list of 146 codes (M2430, M24311, M24312, M24319, M24321, …) would be recognized for^(M24(3|4F?)|S730|T933)(without the$suffix).codebook()takes this a step further. It not only lists the relevant codes but also translates them into plain text. M2430 and M24311 from ICD-10-CM for example would be described as “Pathological dislocation of unspecified joint, not elsewhere classified” and “Pathological dislocation of right shoulder, not elsewhere classified”. A code such as C430 would be recognized by both ICD-10-CM and ICD-10-SE, with the same meaning but in different languages: “Malignant melanoma of lip” and “Malignt melanom på läpp”.
🔗 Use case
Assume we have a list of patients who received surgery at specified dates:
library(coder)
ex_people # Random names and dates included in the package
# A tibble: 100 x 2
name surgery
<chr> <date>
1 Chen, Trevor 2021-01-19
2 Graves, Acineth 2020-10-11
3 Trujillo, Yanelly 2020-09-28
4 Simpson, Kenneth 2020-12-31
5 Chin, Nelson 2020-12-14
6 Le, Christina 2020-07-18
7 Kang, Xuan 2020-10-20
8 Shuemaker, Lauren 2020-07-19
9 Boucher, Teresa 2020-12-25
10 Le, Soraiya 2020-11-29
# ... with 90 more rows
Those patients (among others) were also recorded in a national patient register with date of hospital admissions and corresponding ICD-10 codes. Each patient can have multiple hospital visits over the years. Also, multiple diagnosis might have been recorded at each occasion.
ex_icd10
# A tibble: 2,376 x 4
name admission icd10 hdia
<chr> <date> <chr> <lgl>
1 Tran, Kenneth 2020-08-02 S134A FALSE
2 Tran, Kenneth 2021-01-16 W3319 FALSE
3 Tran, Kenneth 2020-12-26 Y0262 TRUE
4 Tran, Kenneth 2020-11-18 X0488 FALSE
5 Sommerville, Dominic 2021-01-07 V8104 FALSE
6 Sommerville, Dominic 2020-08-18 B853 FALSE
7 Sommerville, Dominic 2021-01-02 Q174 FALSE
8 Sommerville, Dominic 2020-08-23 A227 FALSE
9 Sommerville, Dominic 2020-12-28 H702 FALSE
10 Sommerville, Dominic 2020-04-21 X6051 TRUE
# ... with 2,366 more rows
Using those two data sets, as well as a classification scheme (a classcodes object, here charlson), we can easily identify all Charlson comorbidities for each patient:
ch <-
categorize(
ex_people, # patients of interest
codedata = ex_icd10, # Medical codes from national patient register
cc = "charlson", # Calculate Charlson comorbidity
id = "name", code = "icd10" # Identify column names
)
Classification based on: icd10
Now, how many of those patients were diagnosed with malignancy?
sum(ch$malignancy)
[1] 5
What is the distribution of the combined Charlson comorbidity index for each patient? (This would summarize the general health status of the cohort.)
barplot(table(ch$charlson))
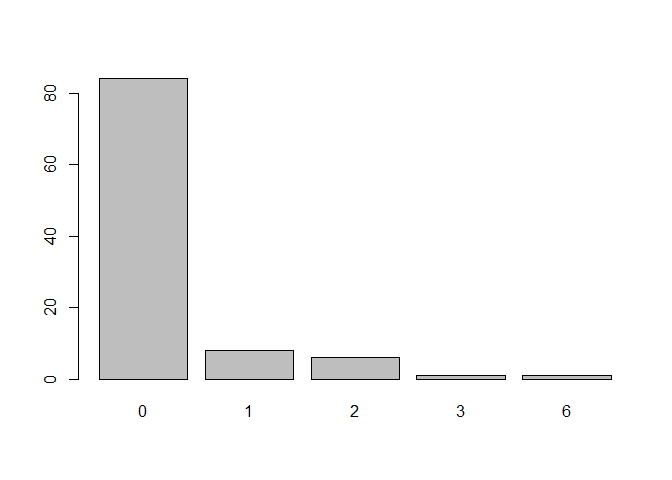
Distribution of the Charlson comorbidity index. Patients in this cohort are healthy. Most have zero comorbidities and only a few have several.
🔗 Further usage
There are many more examples provided at the coder web page. It is easy to constrain the time frame to comorbidities recorded during only one year prior to surgery, or to 90 days for adverse events thereafter, or to use different versions of the regular expressions etc. Core functionality is provided by the categorize() function used above. Alternatively, more fine grained control is provided through a more explicit chain of commands: codify() %>% classify() %>% index().
So far, the package has been used in several research projects (cited in the corresponding JOSS article).
Contributions are most welcome and I would be very happy to hear about possible additional use cases. And please note that there are no hard-coded limitations to patients and/or medical data. The package can be used with all types of codified data!
🔗 Acknowledgement
An early version of the coder package was significantly improved after generous contributions made by Emily C. Zabore and David Robinson during the the peer-review process for rOpenSci. I therefore wish to express my deepest gratitude to both of them, as well as to the rOpenSci community at large!

