Friday, November 18, 2022 From rOpenSci (https://ropensci.org/blog/2022/11/18/ropensci-news-digest-november-2022/). Except where otherwise noted, content on this site is licensed under the CC-BY license.
Dear rOpenSci friends, it’s time for our monthly news roundup!
You can read this post on our blog. Now let’s dive into the activity at and around rOpenSci!
🔗 rOpenSci HQ
🔗 Multilingual Publishing
Open Source and Open Science are global movements, but most of their material and resources are published in English, meaning non-English speakers face a significant barrier to being part of these movements.
Publishing multilingual resources can lower these barriers by increasing access to knowledge, which helps democratize access to quality resources, and therefore increases the potential for contributing to software and open science projects.
We’re excited to announce that with the support of the Chan-Zuckerberg Initiative, NumFOCUS and the R Consortium, we have started translating rOpenSci’s material on best practices for software development, code review, and contribution to open source projects into Spanish. As part of this effort we are also developing guidelines for translating open source resources to a wider audience.
Learn more about our Multilingual Publishing project.
🔗 Champions Program
The application period for our champions program is now closed.
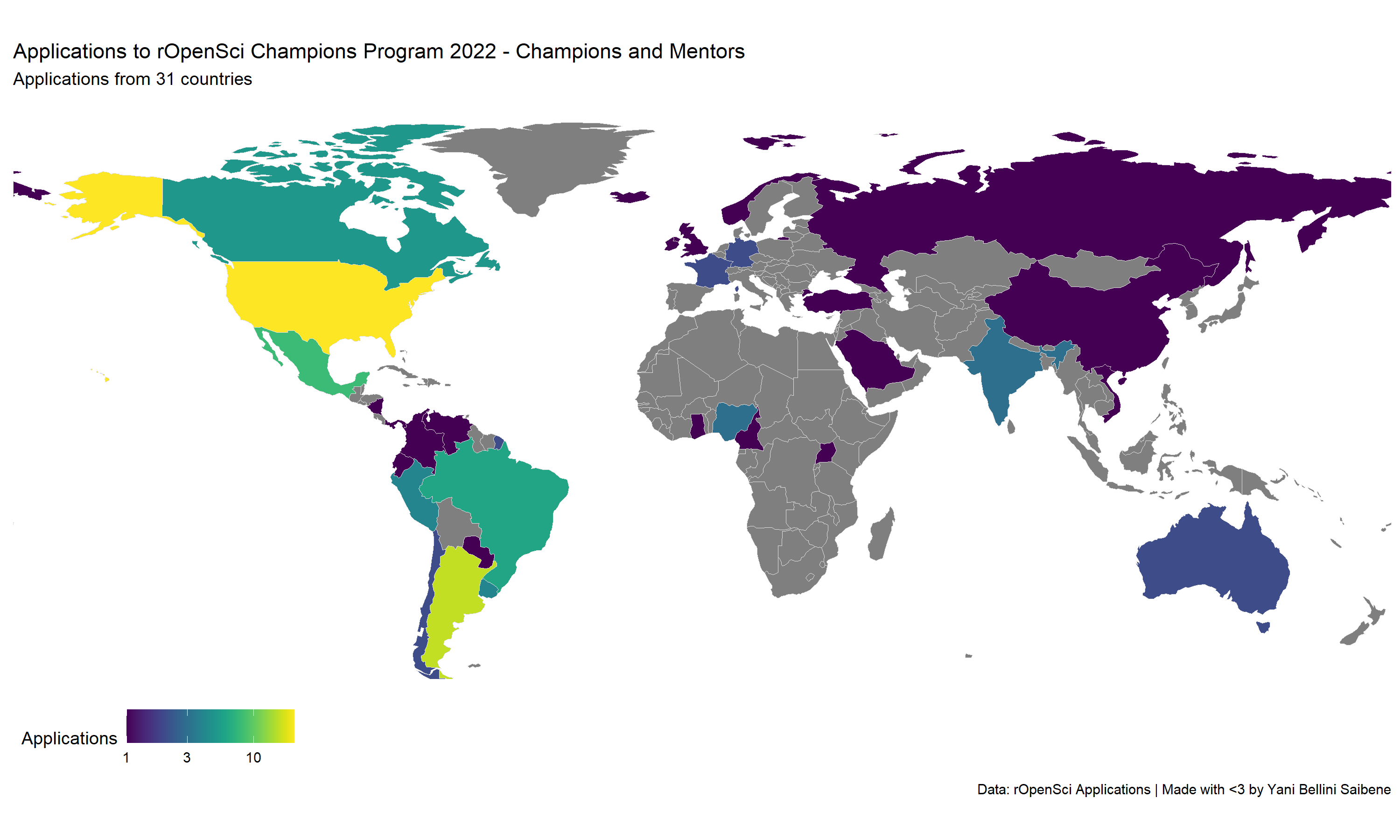
We are very excited about the response from the community! We received 74 applications for Champions and 28 for mentors from 31 countries.
We are very grateful to those who took the time to apply and to all who helped us spread the word about these calls.
Over the next few weeks, we will review the proposals and select the nominees. All applicants will receive feedback on their applications.
Learn more about our Champions Program.
🔗 Coworking sessions continue!
Join us for social coworking & office hours monthly on first Tuesdays! Hosted by Steffi LaZerte and various community hosts. Everyone welcome. No RSVP needed. Consult our Events page to find your local time and how to join.
- Tuesday, December 6th, 9:00 Australian Western / 1:00 UTC “Getting started with targets!” Hosted by community host Nick Tierney and Steffi LaZerte
- Dive into the world of targets! Do some studying; Start setting it up for some of your projects;
- Ask Nick for suggestions how how to get started or tips and tricks.
- Tuesday, Jan 10th, 14:00 European Central / 13:00 UTC “Working with new R users” Hosted by community host Alex Koiter and Steffi LaZerte
- Brainstorm ideas for supporting and encouraging new R users; Annotate a script for a friend or colleague to help them learn;
- Talk to Alex and discuss how to share the love of R with new R users.
- Tuesday, Feb 7th, 9:00 Americas Pacific / 17:00 UTC “Setting up Continuous Integration” Hosted by community host Hugo Gruson and Steffi LaZerte
- Do some reading to learn about Continuous Integration; Setup Continuous Integration on one (or more) of your projects;
- Talk to Hugo and discuss how Continuous Integration can simplify your development process and how to get set up.
And remember, you can always cowork independently on work related to R, work on packages that tend to be neglected, or work on what ever you need to get done!
🔗 rOpenSci communication channels as an alternative to Twitter
Our Twitter account is still active for now but here are alternatives:
🐘 Mastodon account (if you like social media)
🗞️ Newsletter
✍️ Blog
You can read more in our blog post.
🔗 Software 📦
🔗 New packages
The following two packages recently became a part of our software suite:
daiquiri, developed by T. Phuong Quan: Generate reports that enable quick visual review of temporal shifts in record-level data. Time series plots showing aggregated values are automatically created for each data field (column) depending on its contents (e.g. min/max/mean values for numeric data, no. of distinct values for categorical data), as well as overviews for missing values, non-conformant values, and duplicated rows. The resulting reports are shareable and can contribute to forming a transparent record of the entire analysis process. It is designed with Electronic Health Records in mind, but can be used for any type of record-level temporal data (i.e. tabular data where each row represents a single “event”, one column contains the “event date”, and other columns contain any associated values for the event). It is available on CRAN. It has been reviewed by Brad Cannell, and Mauro Lepore.
npi, developed by Frank Farach: Access the United States National Provider Identifier Registry API https://npiregistry.cms.hhs.gov/api/. Obtain and transform administrative data linked to a specific individual or organizational healthcare provider, or perform advanced searches based on provider name, location, type of service, credentials, and other attributes exposed by the API. It is available on CRAN. It has been reviewed by Matthias Grenié, and Emily C. Zabor.
Discover more packages, read more about Software Peer Review.
🔗 New versions
The following twenty packages have had an update since the last newsletter: frictionless (v1.0.2), aorsf (v0.0.4), assertr (v3.0.0), chromer (v0.3), daiquiri (v1.0.1), jagstargets (1.0.4), mctq (v0.3.1), nodbi (v0.9.0), npi (v0.2.0), oai (v0.4.0), rcrossref (v1.2.0), restez (v2.1.3), spiro (v0.1.2), stantargets (0.0.6), stats19 (v2.0.1), stplanr (v1.0.2), tarchetypes (0.7.2), targets (0.14.0), vcr (v1.2.0), and webchem (v1.2.0).
🔗 Software Peer Review
There are thirteen recently closed and active submissions and 2 submissions on hold. Issues are at different stages:
Two at ‘6/approved’:
daiquiri, Data Quality Reporting for Temporal Datasets. Submitted by Phuong Quan.
npi, Access the U.S. National Provider Identifier Registry API. Submitted by Frank Farach.
One at ‘5/awaiting-reviewer(s)-response’:
- phruta, Phylogenetic Reconstruction and Time-dating. Submitted by Cristian Román Palacios.
Four at ‘4/review(s)-in-awaiting-changes’:
hudr, A R interface for accessing HUD (US Department of Housing and Urban Development) APIs. Submitted by Emmet Tam.
octolog, Better Github Action Logging. Submitted by Jacob Wujciak-Jens.
tsbox, Class-Agnostic Time Series. Submitted by Christoph Sax. (Stats).
healthdatacsv, Access data in the healthdata.gov catalog. Submitted by iecastro.
Three at ‘3/reviewer(s)-assigned’:
dynamite, Bayesian Modeling and Causal Inference for Multivariate. Submitted by Santtu Tikka. (Stats).
stochLAB, Stochastic Collision Risk Model. Submitted by Grant. (Stats).
wmm, World Magnetic Model. Submitted by Will Frierson.
One at ‘2/seeking-reviewer(s)’:
- bssm, Bayesian Inference of Non-Linear and Non-Gaussian State Space. Submitted by Jouni Helske. (Stats).
Two at ‘1/editor-checks’:
openalexR, Getting Bibliographic Records from OpenAlex Database Using DSL. Submitted by Trang Le.
dfms, Dynamic Factor Models. Submitted by Sebastian Krantz.
Find out more about Software Peer Review and how to get involved.
🔗 On the blog
Become a Mentor for rOpenSci Champions! by Yanina Bellini Saibene. rOpenSci is seeking mentors to support our inaugural cohort of rOpenSci Champions. Could you offer insight and advice to our selected Champions? Learn the details and express your interest.
rOpenSci’s Communication Channels: Twitter by Yanina Bellini Saibene, and Steffi LaZerte. We announce our actions on rOpenSci’s communication channels as alternatives to Twitter.
Canales de comunicación de rOpenSci: Twitter by Yanina Bellini Saibene, and Steffi LaZerte. Anunciamos nuestras acciones en los canales de comunicación de rOpenSci como alternativas a Twitter.
🔗 Use cases
One use case of our packages and resources has been reported since we sent the last newsletter.
- Adding missing EXIF data to wildlife trail camera images. Reported by Neil Saunders.
Explore other use cases and report your own!
🔗 Call for package (co-)maintainers
🔗 Call for maintainers
There are still a few packages to adopt from our recent blog post. To volunteer, comment in the corresponding volunteering issue. Thank you!
- wikitaxa, Taxonomic Information from ‘Wikipedia’. ‘Taxonomic’ information from ‘Wikipedia’, ‘Wikicommons’, ‘Wikispecies’, and ‘Wikidata’. Functions included for getting taxonomic information from each of the sources just listed, as well performing taxonomic search. Issue for volunteering.
- rgnparser, Parse Scientific Names. Parse scientific names using ‘gnparser’, written in Go. ‘gnparser’ parses scientific names into their component parts; it utilizes a Parsing Expression Grammar specifically for scientific names. Issue for volunteering.
- RSelenium, R Bindings for ‘Selenium WebDriver’. Provides a set of R bindings for the ‘Selenium 2.0 WebDriver’ using the ‘JsonWireProtocol’. ‘Selenium 2.0 WebDriver’ allows driving a web browser natively as a user would either locally or on a remote machine using the Selenium server it marks a leap forward in terms of web browser automation. Issue for volunteering.
- elastic, General Purpose Interface to ‘Elasticsearch’. Connect to ‘Elasticsearch’, a ‘NoSQL’ database built on the ‘Java’ Virtual Machine. Interacts with the ‘Elasticsearch’ ‘HTTP’ API, including functions for setting connection details to ‘Elasticsearch’ instances, loading bulk data, searching for documents with both ‘HTTP’ query variables and ‘JSON’ based body requests. Issue for volunteering.
- Rclean, A Tool for Writing Cleaner, More Transparent Code. To create clearer, more concise code provides this toolbox helps coders to isolate the essential parts of a script that produces a chosen result, such as an object, tables and figures written to disk. Issue for volunteering.
🔗 Call for comaintainers
Refer to our recent blog post to identify packages where help is especially wished for!
🔗 Package development corner
Some useful tips for R package developers. 👀
🔗 Tired of typing #’ before function examples?
You can write your function examples in separate scripts and then refer to them using the roxygen2 @example (no s!) tag.
You’d write
#' @example man/examples/foo.R
and in man/examples/foo.R
# basic usage of foo
foo(basic = TRUE)
# elaborate usage of foo
foo(basic = FALSE)
Downsides of using this are that it might surprise contributors, and that someone who’d look for the source of the example through, say, the link indicated on a pkgdown reference page, would not get to the example source directly.
Thanks to Hugo Gruson for reminding this in the rOpenSci semi-open slack.
🔗 Display a message or warning only once per session?
If that’s your need, know that rlang::warn() and rlang::inform() have a handy .frequency argument, as reported by Jon Harmon on Posit Community forum.
🔗 Run some tests on continuous integration only?
Say you have some slow and fragile tests querying an API. If you want to run them on continuous integration only, refer to Bryce Mecum’s blog post.
🔗 How to use additional packages for the pkgdown website only
Say a package is needed for a pkgdown article of your package (but not a vignette), or for nicer autolinking of a reference to a function (for instance if you recommend usethis::create_package()).
Assuming you’re building your website with GitHub Actions from r-lib/actions (that you might have gotten via usethis).
If so, you can use the Config/Needs/website field in DESCRIPTION.
Here is an example, in pkgdown itself
Config/Needs/website: usethis, servr
The idea of custom fields is mentioned in the second edition of the R packages book.
Note that it works for rOpenSci packages, whose documentation websites are built with R-universe!
🔗 Last words
Thanks for reading! If you want to get involved with rOpenSci, check out our Contributing Guide that can help direct you to the right place, whether you want to make code contributions, non-code contributions, or contribute in other ways like sharing use cases.
If you haven’t subscribed to our newsletter yet, you can do so via a form. Until it’s time for our next newsletter, you can keep in touch with us via our website and Twitter account.

