Thursday, April 2, 2026 From rOpenSci (https://ropensci.org/blog/2026/04/02/tree-sitter-overview/). Except where otherwise noted, content on this site is licensed under the CC-BY license.
A little bit less than two years ago, building on work by Jim Hester and Kevin Ushey, Davis Vaughan completed a very impactful JavaScript file for the R community: an R grammar for the Tree-sitter parsing generator. He even got a round of applause for it during a talk at the useR! 2024 conference! So, did he get cheered for… grammatical rules in a JavaScript file? 😅
No, the audience was excited about the improved developer experience for R that this file unlocked. R tooling around Tree-sitter is how you get
- reformatting through Air and linting through Jarl;
- auto-completion or help on hover in the Positron IDE;
- better search for R on GitHub;
- and more!
In this post, we’ll explain what Tree-sitter is, and how tools built on Tree-sitter can benefit your R development workflow.
🔗 Code parsing: what is Tree-sitter?
Tree-sitter is a code parsing generator written in C, with bindings existing in several languages including Rust (and R!).
Let’s rewind a little bit. What does it mean to parse code?
Basically, given a string of code like
a <- mean(x, na.rm = TRUE)
How do you know that mean is a function name, na.rm an argument name, TRUE a logical? You have to parse that code into what’s called a parse tree. You do that in your head when reading R code. 😸
R itself can obviously parse R code, thanks to its grammar. See for instance the commit that introduced R’s native pipe, which necessitated extending R’s syntax thus modifying its grammar.
You can use parse() and getParseData() to parse R code.
parse(
text = "a <- mean(x, na.rm = TRUE)",
keep.source = TRUE
) |>
getParseData()
#> line1 col1 line2 col2 id parent token terminal text
#> 23 1 1 1 26 23 0 expr FALSE
#> 1 1 1 1 1 1 3 SYMBOL TRUE a
#> 3 1 1 1 1 3 23 expr FALSE
#> 2 1 3 1 4 2 23 LEFT_ASSIGN TRUE <-
#> 21 1 6 1 26 21 23 expr FALSE
#> 4 1 6 1 9 4 6 SYMBOL_FUNCTION_CALL TRUE mean
#> 6 1 6 1 9 6 21 expr FALSE
#> 5 1 10 1 10 5 21 '(' TRUE (
#> 7 1 11 1 11 7 9 SYMBOL TRUE x
#> 9 1 11 1 11 9 21 expr FALSE
#> 8 1 12 1 12 8 21 ',' TRUE ,
#> 13 1 14 1 18 13 21 SYMBOL_SUB TRUE na.rm
#> 14 1 20 1 20 14 21 EQ_SUB TRUE =
#> 15 1 22 1 25 15 16 NUM_CONST TRUE TRUE
#> 16 1 22 1 25 16 21 expr FALSE
#> 17 1 26 1 26 17 21 ')' TRUE )
Or you could transform that same data into XML using Gábor Csárdi’s {xmlparsedata}:
parse(
text = "a <- mean(x, na.rm = TRUE)",
keep.source = TRUE
) |>
xmlparsedata::xml_parse_data(pretty = TRUE) |>
xml2::read_xml() |>
as.character() |>
cat()
#> <?xml version="1.0" encoding="UTF-8" standalone="yes"?>
#> <exprlist>
#> <expr line1="1" col1="1" line2="1" col2="26" start="28" end="53">
#> <expr line1="1" col1="1" line2="1" col2="1" start="28" end="28">
#> <SYMBOL line1="1" col1="1" line2="1" col2="1" start="28" end="28">a</SYMBOL>
#> </expr>
#> <LEFT_ASSIGN line1="1" col1="3" line2="1" col2="4" start="30" end="31"><-</LEFT_ASSIGN>
#> <expr line1="1" col1="6" line2="1" col2="26" start="33" end="53">
#> <expr line1="1" col1="6" line2="1" col2="9" start="33" end="36">
#> <SYMBOL_FUNCTION_CALL line1="1" col1="6" line2="1" col2="9" start="33" end="36">mean</SYMBOL_FUNCTION_CALL>
#> </expr>
#> <OP-LEFT-PAREN line1="1" col1="10" line2="1" col2="10" start="37" end="37">(</OP-LEFT-PAREN>
#> <expr line1="1" col1="11" line2="1" col2="11" start="38" end="38">
#> <SYMBOL line1="1" col1="11" line2="1" col2="11" start="38" end="38">x</SYMBOL>
#> </expr>
#> <OP-COMMA line1="1" col1="12" line2="1" col2="12" start="39" end="39">,</OP-COMMA>
#> <SYMBOL_SUB line1="1" col1="14" line2="1" col2="18" start="41" end="45">na.rm</SYMBOL_SUB>
#> <EQ_SUB line1="1" col1="20" line2="1" col2="20" start="47" end="47">=</EQ_SUB>
#> <expr line1="1" col1="22" line2="1" col2="25" start="49" end="52">
#> <NUM_CONST line1="1" col1="22" line2="1" col2="25" start="49" end="52">TRUE</NUM_CONST>
#> </expr>
#> <OP-RIGHT-PAREN line1="1" col1="26" line2="1" col2="26" start="53" end="53">)</OP-RIGHT-PAREN>
#> </expr>
#> </expr>
#> </exprlist>
In both cases, you recognize words such as LEFT_ASSIGN or SYMBOL_FUNCTION_CALL. Parsing is an essential step before the code is actually executed, but parsed code can also be used for other purposes, such as analyzing code without brittle regular expressions (does it call a particular function?), navigating code (going from a function call to the definition of that function), or modifying code (replacing all occurrences of a function with another one).
Now, Tree-sitter performs this same code parsing but faster especially thanks to its support of incremental parsing – which is key to updating the syntax tree as you are typing in your editor for instance! Tree-sitter is agnostic in that it can parse any code as long as there is a grammar for it (think, Rosetta Stone plugins). It’s been used for many languages which means many tools have been built around it.
To have Tree-sitter “learn” a new language you need to give it a file containing the definition of the syntax of that language, what’s called a grammar. This is where the aforementioned JavaScript file by Davis Vaughan and collaborators comes into play! The treesitter-r repo, which provides a translation of the R grammar in the format expected by Tree-sitter, is the base of all tools presented in this post which use R code as their input.
Here’s how to use the {treesitter} R package for the same code as earlier. The {treesitter} R package allows us to use Tree-sitter from R. To parse R code with it, we need the language() function from {treesitter.r}1.
library(treesitter)
#>
#> Attaching package: 'treesitter'
#> The following object is masked from 'package:base':
#>
#> range
language <- treesitter.r::language()
parser <- parser(language)
text <- "a <- mean(x, na.rm = TRUE)"
parser_parse(parser, text)
#> <tree_sitter_tree>
#>
#> ── Text ────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────
#> a <- mean(x, na.rm = TRUE)
#>
#> ── S-Expression ────────────────────────────────────────────────────────────────────────────────────────────────────────────────
#> (program [(0, 0), (0, 26)]
#> (binary_operator [(0, 0), (0, 26)]
#> lhs: (identifier [(0, 0), (0, 1)])
#> operator: "<-" [(0, 2), (0, 4)]
#> rhs: (call [(0, 5), (0, 26)]
#> function: (identifier [(0, 5), (0, 9)])
#> arguments: (arguments [(0, 9), (0, 26)]
#> open: "(" [(0, 9), (0, 10)]
#> argument: (argument [(0, 10), (0, 11)]
#> value: (identifier [(0, 10), (0, 11)])
#> )
#> (comma [(0, 11), (0, 12)])
#> argument: (argument [(0, 13), (0, 25)]
#> name: (identifier [(0, 13), (0, 18)])
#> "=" [(0, 19), (0, 20)]
#> value: (true [(0, 21), (0, 25)])
#> )
#> close: ")" [(0, 25), (0, 26)]
#> )
#> )
#> )
#> )
Tree-sitter is the workhorse of many tools, that are mentioned in the diagram below. All of them are dependent on tree-sitter and the R grammar provided to it. Some of them are command-line interfaces (CLIs), while others are R packages.
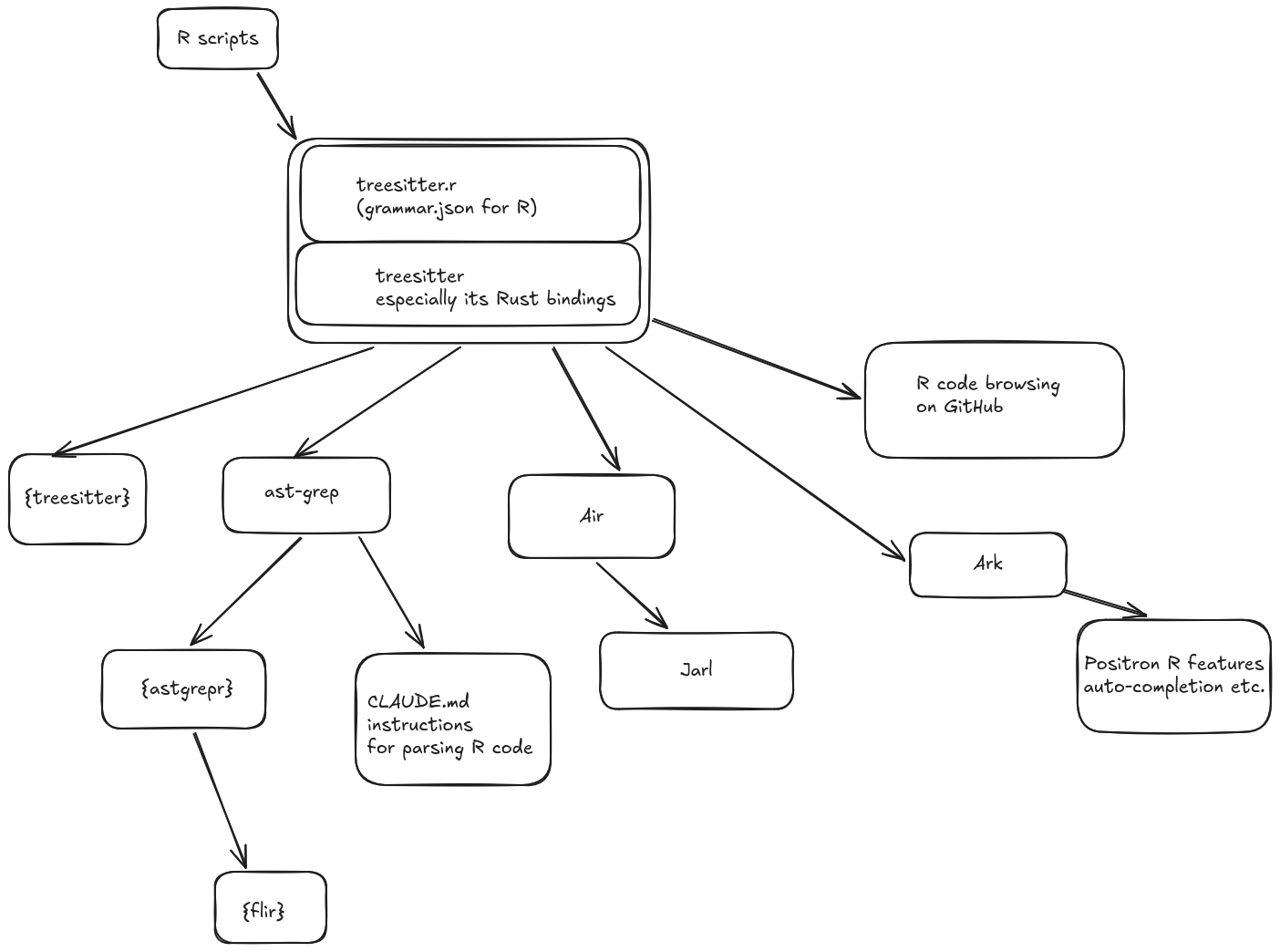
🔗 Browsing code interactively: Positron IDE, GitHub
The real reason why the audience applauded Davis Vaughan is that he explained how the R grammar for Tree-sitter had been deployed to GitHub so that we get almost as good experience browsing R code on GitHub as browsing, say, JS code. If we search for a function name in a repository for instance, its definition will be indicated in the search results. See Davis’ slides (also available in PDF), or refer to the video below showing how typing vetiver_model in the search bar from the R vetiver repo makes the function definition the first result, on which one can click to land into the definition.
Also very useful is the use of Tree-sitter by Ark, the R kernel used in the Positron IDE. Ark is how you get autocompletion and help on hover in Positron. The video below shows how you can extend the selection to further steps of a pipeline in Positron.
This use case of Tree-sitter is also featured in Davis’ slides. See also Lionel Henry’s and Davis Vaughan’s talk about Ark at posit conf 2024, especially the part about code assistance.
Other development environments such as Emacs also have support for Tree-sitter.
🔗 Searching/browsing code
You can parse and search R code using the {treesitter} R package and treesitter query syntax. The {treesitter} R package is a dependency of the {gander} package by Simon Couch, that is meant to be used for a better experience with LLMs when writing R code. Another use case of the {treesitter} R package is the {igraph.r2cdocs} extension to {roxygen2} for the {igraph} package, that parses all of igraph R code to then be able to identify, for each exported function, whether it (in)directly calls a function whose name ends with _impl, indicating a wrapper to a C igraph function whose docs can be then be linked from the manual of the R function.
The {pkgdepends} package calls Tree-sitter (C) to detect dependencies in files. Below we run it on the source of the saperlipopette R package.
pkgdepends::scan_deps(
"../../../../../CHAMPIONS/saperlipopette",
"../../../../../CHAMPIONS"
)
#>
#> Dependencies:
#> + brio @ R/blame.R, R/check-editor.R, R/clean-dir.R, R/committed-to-main.R, R/committed-to-wrong-branch.R, R/conflict…
#> + cli @ inst/exo_bisect-Rprofile.en.R, inst/exo_bisect-Rprofile.es.R, inst/exo_bisect-Rprofile.fr.R, inst/exo_blame-…
#> + devtools @ saperlipopette.Rproj
#> + fs @ R/blame.R, R/check-editor.R, R/clean-dir.R, R/committed-to-main.R, R/committed-to-wrong-branch.R, R/conflict…
#> + gert @ inst/exo_check_editor-Rprofile.en.R, inst/exo_check_editor-Rprofile.es.R, inst/exo_check_editor-Rprofile.fr.…
#> + knitr @ README.Rmd
#> + parsedate @ R/utils-git.R
#> + purrr @ R/create-all.R, R/debug.R, R/log-deleted-file.R, R/log-deleted-line.R, R/revparse.R, R/roxygen2.R, R/worktre…
#> + rlang @ R/create-all.R, R/roxygen2.R, R/utils-fs.R, R/utils-usethis.R, R/zzz.R
#> + rmarkdown @ README.Rmd, vignettes/saperlipopette.qmd
#> + roxygen2 @ R/roxygen2.R, saperlipopette.Rproj
#> + saperlipopette @ README.Rmd, vignettes/saperlipopette.qmd
#> + tibble @ R/roxygen2.R
#> + usethis @ R/blame.R, R/check-editor.R, R/clean-dir.R, R/committed-to-main.R, R/committed-to-wrong-branch.R, R/conflict…
#> + vctrs @ R/roxygen2.R
#> + withr @ R/blame.R, R/check-editor.R, R/clean-dir.R, R/committed-to-main.R, R/committed-to-wrong-branch.R, R/conflict…
#>
#> Test dependencies:
#> + fs @ tests/testthat/test-blame.R, tests/testthat/test-check-editor.R, tests/testthat/test-clean-dir.R, tests/test…
#> + gert @ tests/testthat/test-blame.R, tests/testthat/test-clean-dir.R, tests/testthat/test-committed-to-main.R, tests…
#> + rlang @ tests/testthat/test-blame.R, tests/testthat/test-check-editor.R, tests/testthat/test-clean-dir.R, tests/test…
#> + saperlipopette @ tests/testthat.R
#> + testthat @ tests/testthat.R
#> + withr @ tests/testthat/test-blame.R, tests/testthat/test-check-editor.R, tests/testthat/test-clean-dir.R, tests/test…
ast-grep is a useful tool built on Tree-sitter for searching and re-writing code, with a clearer query syntax than Tree-sitter’s. Its name is reminiscent of grep, but with ast-grep we do not need to write brittle regular expressions 😸. {astgrepr} by Etienne Bacher is an R wrapper to the Rust bindings of ast-grep, and is used in Etienne’s {flir} package for refactoring code.
The ast-grep command-line interface (CLI) itself is featured in a useful blog post by Emil Hvitfeldt where he explains how to document the usage of ast-grep for Claude.
🔗 Formatting and linting: Air, Jarl
Speaking of CLIs…

Air, by Davis Vaughan and Lionel Henry, is a CLI built on Tree-sitter, in Rust. It reformats code blazingly fast.
Jarl, by Etienne Bacher, is a CLI built on Air, therefore also on Tree-sitter, in Rust. It lints and fixes code, also blazingly fast. It can even detect unreachable code, unused functions and duplicated function definitions.
In both of these examples, the creation of CLIs wrapping Rust bindings was more efficient than the creation of R packages wrapping the {treesitter} R package, for several reasons:
- Rust CLIs can edit code very fast2;
- CLIs are integrated in extensions for popular IDEs (for instance Positron);
- a CLI is easier to install on CI than an R package that needs, well, an R installation.
🔗 More tools
A brief mention of some other interesting tools we’ve explored a bit less.
🔗 Configuring: {ts} for parsing JSON and TOML (not R!)
The {ts} package by Gábor Csárdi is the backbone of two R packages used for editing and manipulating:
Compared to existing parsers in R for those formats, these two packages preserve comments.
🔗 Testing code: {muttest}
Mutation testing is a kind of testing where you, say, randomly swap + with - in your code (you mutate it) and you run your tests to see whether they catch the mutant. The {muttest} package by Jakub Sobolewski is an R package for mutation testing, that depends on the {treesitter} R package.
🔗 Diffing code: difftastic
The difftastic CLI by Wilfred Hughes is “a structural diff tool that understands syntax”. ✨ This means that difftastic doesn’t only compare line or “words” but actual syntax by looking at lines around the lines that changed (by default, 3). Even better, it understands R out of the box. See this blog post with examples of R code diffing.
🔗 Conclusion: more to come?
In this post, we’ve presented an overview of Tree-sitter based tooling for R or in R.
Note that this ecosystem of tools is very actively developed, so some tools might come and go. However, the idea that plugging the R grammar into a general parsing generator brings cool features to us R developers, will remain true. Maybe you will contribute to this ecosystem, either through an existing tool or by creating a new one?
We could also parse C code with it using {treesitter.c}. ↩︎
Rust is a lower level language than R so has less overhead; furthermore this kind of Rust code can be easily parallelized. ↩︎

