Wednesday, March 1, 2017 From rOpenSci (https://ropensci.org/blog/2017/03/01/ccafs-release/). Except where otherwise noted, content on this site is licensed under the CC-BY license.
I’ve recently released the new package ccafs, which provides access to data from Climate Change, Agriculture and Food Security (CCAFS; http://ccafs-climate.org/) General Circulation Models (GCM) data. GCM’s are a particular type of climate model, used for weather forecasting, and climate change forecasting - read more at https://en.wikipedia.org/wiki/General_circulation_model.
ccafs falls in the data client camp - its focus is on getting users
data - many rOpenSci packages
fall into this area. These kinds of packages are important so that
scientists don’t have to recreate the wheel themselves every time, but
instead use one client that everyone else uses.
CCAFS GCM data files are .zip files with a bunch of files inside. The
individual files are in ARC ASCII format (https://en.wikipedia.org/wiki/Esri_grid#ASCII) -
a plain text data format, but still require painful manipulation/wrangling to
get into an easily consumable format. The files have a .asc file extension.
For each .asc file, the first 6 lines of each file indicate the reference of
the grid (number of columns and rows, corner coordinates, cellsize, and missing
data value), followed by the actual data values, delimited with single
space characters.
There’s a related binary format - but its proprietary, so nevermind.
The workflow with ccafs for most users will likely be as follows:
- Search for data they want:
cc_search() - Fetch/download data:
cc_data_fetch() - Reaad data:
cc_data_read()
I’ll dive into more details below.
🔗 Installation
First, install the package.
install.packages("ccafs")
Then load ccafs
library("ccafs")
🔗 Search for data
Searching CCAF’s data holdings is not as easy as it could be as they don’t provide any programmatic way to do so. However, we provide a way to search using their web interface from R.
You can search by the numbers representing each possible value for
each parameter. See the ?'ccafs-search' for help on what the numbers
refer to.
(result1 <- cc_search(file_set = 4, scenario = 6, model = 2, extent = "global",
format = "ascii", period = 5, variable = 2, resolution = 3))
#> [1] "http://gisweb.ciat.cgiar.org/ccafs_climate/files/data/ipcc_4ar_ciat/sres_b1/2040s/bccr_bcm2_0/5min/bccr_bcm2_0_sres_b1_2040s_prec_5min_no_tile_asc.zip"
Alternatively, you can use the helper list cc_params where you can reference
options by name; the downside is that this leads to very verbose code.
(result2 <- cc_search(file_set = cc_params$file_set$`Delta method IPCC AR4`,
scenario = cc_params$scenario$`SRES B1`,
model = cc_params$model$bccr_bcm2_0,
extent = cc_params$extent$global,
format = cc_params$format$ascii,
period = cc_params$period$`2040s`,
variable = cc_params$variable$Precipitation,
resolution = cc_params$resolution$`5 minutes`))
#> [1] "http://gisweb.ciat.cgiar.org/ccafs_climate/files/data/ipcc_4ar_ciat/sres_b1/2040s/bccr_bcm2_0/5min/bccr_bcm2_0_sres_b1_2040s_prec_5min_no_tile_asc.zip"
If you already know what you want in terms of file paths, you can query
Amazon S3 directly with cc_list_keys() (the data file come from Amazon S3):
cc_list_keys(max = 3)
#> # A tibble: 3 × 5
#> Key LastModified
#> <chr> <chr>
#> 1 ccafs/ 2014-02-28T15:15:45.000Z
#> 2 ccafs/2014-05-24-01-19-33-3A0DFF1F86F3E7F7.txt 2014-07-01T02:15:51.000Z
#> 3 ccafs/amzn.csv 2014-02-28T15:21:32.000Z
#> # ... with 3 more variables: ETag <chr>, Size <chr>, StorageClass <chr>
When using cc_list_keys(), you’ll get not just .zip files that can be
downloaded, but also directories. So beware that if you’re going after grabbing
“keys” for files that can be downloaded, you’re looking for .zip files.
🔗 Fetch and read data
Once you get links from cc_search() or “keys” from cc_list_keys(), you
can pass either to cc_data_fetch() - which normalizes the input - so it
doesn’t matter whether you pass in e.g.,
http://gisweb.ciat.cgiar.org/ccafs_climate/files/data/ipcc_4ar_ciat/
sres_b1/2040s/bccr_bcm2_0/5min/bccr_bcm2_0_sres_b1_2040s_prec_5min_no_tile_asc.zip
or
ccafs_climate/files/data/ipcc_4ar_ciat/sres_b1/2040s/bccr_bcm2_0/5min/
bccr_bcm2_0_sres_b1_2040s_prec_5min_no_tile_asc.zip
Let’s download data with cc_data_fetch() using the result we got above
using cc_search():
xx <- cc_data_fetch(result2)
Then we can read data with cc_data_read():
(dat <- cc_data_read(xx))
#> class : RasterStack
#> dimensions : 1800, 4320, 7776000, 12 (nrow, ncol, ncell, nlayers)
#> resolution : 0.08333333, 0.08333333 (x, y)
#> extent : -180, 180, -60, 90 (xmin, xmax, ymin, ymax)
#> coord. ref. : NA
#> names : prec_1, prec_10, prec_11, prec_12, prec_2, prec_3, prec_4, prec_5, prec_6, prec_7, prec_8, prec_9
#> min values : -2147483648, -2147483648, -2147483648, -2147483648, -2147483648, -2147483648, -2147483648, -2147483648, -2147483648, -2147483648, -2147483648, -2147483648
#> max values : 2147483647, 2147483647, 2147483647, 2147483647, 2147483647, 2147483647, 2147483647, 2147483647, 2147483647, 2147483647, 2147483647, 2147483647
Which gives a raster class object, which you are likely familiar with - which
opens up all the tools that deal with raster class objects, yay!
You can easily plot the data with the plot method from the raster package.
library("raster")
plot(dat)
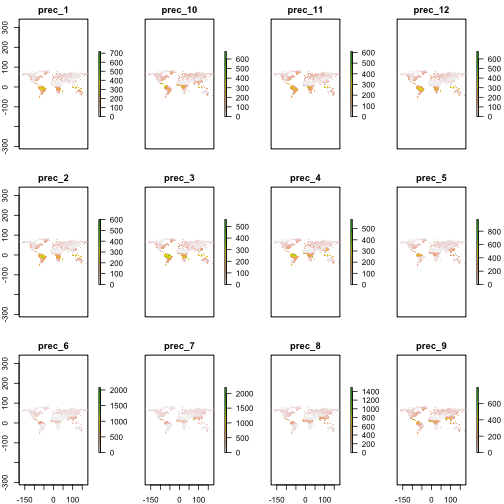
visualize CCAFS data
🔗 Caching
For a better user experience, we cache files for you. That means when we download data, we put the files in a known location. When a user tries to download the same data again, we look to see if it’s already been downloaded, and use the cached version - if we don’t have it already, we download it.
Of course, CCAFS may change their files, so you may not want the cached version, but the new version from them. We provide tools to inspect your cached files, and delete them.
List your cached files:
cc_cache_list()
#> [1] "/Users/sacmac/Library/Caches/ccafs/bcc_csm1_1_m_rcp2_6_2030s_bio_10min_r1i1p1_no_tile_asc"
#> [2] "/Users/sacmac/Library/Caches/ccafs/bcc_csm1_1_m_rcp2_6_2030s_bio_10min_r1i1p1_no_tile_asc.zip"
#> [3] "/Users/sacmac/Library/Caches/ccafs/bcc_csm1_1_m_rcp2_6_2030s_bio_10min_r1i1p1_no_tile_asc/bio_1.asc"
#> [4] "/Users/sacmac/Library/Caches/ccafs/bcc_csm1_1_m_rcp2_6_2030s_bio_10min_r1i1p1_no_tile_asc/bio_10.asc"
#> [5] "/Users/sacmac/Library/Caches/ccafs/bcc_csm1_1_m_rcp2_6_2030s_bio_10min_r1i1p1_no_tile_asc/bio_11.asc"
#> [6] "/Users/sacmac/Library/Caches/ccafs/bcc_csm1_1_m_rcp2_6_2030s_bio_10min_r1i1p1_no_tile_asc/bio_12.asc"
#> [7] "/Users/sacmac/Library/Caches/ccafs/bcc_csm1_1_m_rcp2_6_2030s_bio_10min_r1i1p1_no_tile_asc/bio_13.asc"
#> [8] "/Users/sacmac/Library/Caches/ccafs/bcc_csm1_1_m_rcp2_6_2030s_bio_10min_r1i1p1_no_tile_asc/bio_14.asc"
#> [9] "/Users/sacmac/Library/Caches/ccafs/bcc_csm1_1_m_rcp2_6_2030s_bio_10min_r1i1p1_no_tile_asc/bio_15.asc"
#> [10] "/Users/sacmac/Library/Caches/ccafs/bcc_csm1_1_m_rcp2_6_2030s_bio_10min_r1i1p1_no_tile_asc/bio_16.asc"
...
Get details on all files or a specific file:
# cc_cache_details() # details for all files
cc_cache_details(cc_cache_list()[1])
#> <ccafs cached files>
#> directory: /Users/sacmac/Library/Caches/ccafs
#>
#> file: /bcc_csm1_1_m_rcp2_6_2030s_bio_10min_r1i1p1_no_tile_asc
#> size: 0.001 mb
Be careful with cc_cache_delete_all() as you will delete all your cached
files.
🔗 ccafs software review
I want to touch briefly on the software review for this package. The reviews
for ccafs were great, and I think the package was greatly improved via the
review process.
Michael Koontz and Manuel Ramon
did reviews for ccafs.
One thing in particular that improved about ccafs was the user interface -
that is, the programmatic interface. One feature about the interface was
adding the cc_search() function. When I started developing ccafs, I didn’t
see a way to programmatically search CCAFS data - other than the Amazon S3
data, which isn’t really search, but more like listing files in a directory -
so I just left it at that. During the reviews, reviewers wanted a clear workflow
for potential users - the package as submitted for review didn’t really have a
clear workflow; it was
- Know what you want already (
cc_list_keyshelped get real paths at least) - Download data
- Read data
Which is not ideal. There should be a discovery portion to the workflow. So I decided to dig into possibly querying the CCAFS web portal itself. That panned out, and the workflow we have now is much better:
- Search for data with all the same variables you would on the CCAFS website
- Download data
- Read data
This is much better!
As always, reviews improved the documentation a lot by pointing out areas that could use improvement - which all users will greatly benefit from.
A new vignette (https://cran.rstudio.com/web/packages/ccafs/vignettes/amazon_s3_keys.html) was added in the review process to explain how to get a “key”, a URL for CCAFS data.
🔗 To Do and Feedback
There’s probably lots of improvements that can be made - I’m looking forward to getting feedback from users on any bugs or feature requests. One immediate thing is to make the cache details more compact.

